import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
CSV_URL = ("https://raw.githubusercontent.com/livingphysics/bioreactor_v3/"
"main/src/bioreactor_data/bioreactor_data.csv")
df = pd.read_csv(CSV_URL)
times = df['elapsed_time'].values.astype(float)
measurements = df['Eyespy_sct_V'].values.astype(float)
pump_cum = df['pump_inflow_time_s'].values.astype(float)
pump_events = np.zeros(len(times), dtype=bool)
pump_events[1:] = np.diff(pump_cum) > 0
def run_ekf_replay(times, measurements, pump_events,
R=0.001, Q_growth_rate=5e-12,
initial_growth_rate=1.0, initial_P_r=0.0005**2,
pump_distrust_cycles=10, pump_distrust_P_od=None):
n = len(times)
if pump_distrust_P_od is None:
pump_distrust_P_od = 10.0 * R
od_est = np.full(n, np.nan); growth_rate = np.full(n, np.nan)
doubling_time_s = np.full(n, np.nan); od_std = np.full(n, np.nan)
r_std = np.full(n, np.nan); dt_std = np.full(n, np.nan)
x = np.array([measurements[0], initial_growth_rate])
P = np.array([[R, 0.0], [0.0, initial_P_r]])
distrust_counter = 0
last_time = times[0]
dt_median = np.median(np.diff(times))
od_est[0] = x[0]; growth_rate[0] = x[1]
od_std[0] = np.sqrt(R); r_std[0] = np.sqrt(initial_P_r)
I2 = np.eye(2); H_vec = np.array([1.0, 0.0])
for i in range(1, n):
z_k = measurements[i]
if np.isnan(z_k):
od_est[i] = od_est[i-1]; growth_rate[i] = growth_rate[i-1]
doubling_time_s[i] = doubling_time_s[i-1]
od_std[i] = od_std[i-1]; r_std[i] = r_std[i-1]; dt_std[i] = dt_std[i-1]
continue
last_time = times[i]
if pump_events[i]:
distrust_counter = pump_distrust_cycles
od_k, r_k = x
x_pred = np.array([od_k * r_k, r_k])
F = np.array([[r_k, od_k], [0.0, 1.0]])
Q_mat = np.array([[0.0, 0.0], [0.0, Q_growth_rate]])
P_pred = F @ P @ F.T + Q_mat
if pump_events[i] or distrust_counter > 0:
P_pred[0, 0] = pump_distrust_P_od
P_pred[0, 1] = 0.0; P_pred[1, 0] = 0.0
x_pred[0] = z_k
if not pump_events[i]:
distrust_counter -= 1
y = z_k - x_pred[0]
S = P_pred[0, 0] + R
K = P_pred[:, 0] / S
x_updated = x_pred + K * y
P_updated = (I2 - np.outer(K, H_vec)) @ P_pred
if abs(z_k - x_pred[0]) > 5.0 * np.sqrt(R):
P_updated[0, 1] = 0.0; P_updated[1, 0] = 0.0
P_updated[0, 0] = (x_updated[0] - z_k) ** 2
x, P = x_updated, P_updated
od_est[i] = x[0]; growth_rate[i] = x[1]
od_std[i] = np.sqrt(P[0, 0]); r_std[i] = np.sqrt(P[1, 1])
r_est = x[1]
if r_est > 1.0:
ln_r = np.log(r_est)
dt_val = dt_median * np.log(2.0) / ln_r
if dt_val <= 86400.0:
doubling_time_s[i] = dt_val
dt_std[i] = dt_median * np.log(2.0) * r_std[i] / (r_est * ln_r ** 2)
return dict(od_est=od_est, growth_rate=growth_rate,
doubling_time_s=doubling_time_s,
od_std=od_std, r_std=r_std, dt_std=dt_std)
result = run_ekf_replay(times, measurements, pump_events, Q_growth_rate=5e-12)
t_h = times / 3600.0
mask = t_h <= 9.0
t_h = t_h[mask]
raw_od = measurements[mask]
ekf_od = result['od_est'][mask]
od_sd = result['od_std'][mask]
dt_min = result['doubling_time_s'][mask] / 60.0
dt_sd_min = result['dt_std'][mask] / 60.0
temp = df['temperature_C'].values.astype(float)[mask]
pump_times = t_h[pump_events[mask]]
fig, (ax_od, ax_dt, ax_temp) = plt.subplots(3, 1, figsize=(11, 7.5), sharex=True)
fig.subplots_adjust(hspace=0.12)
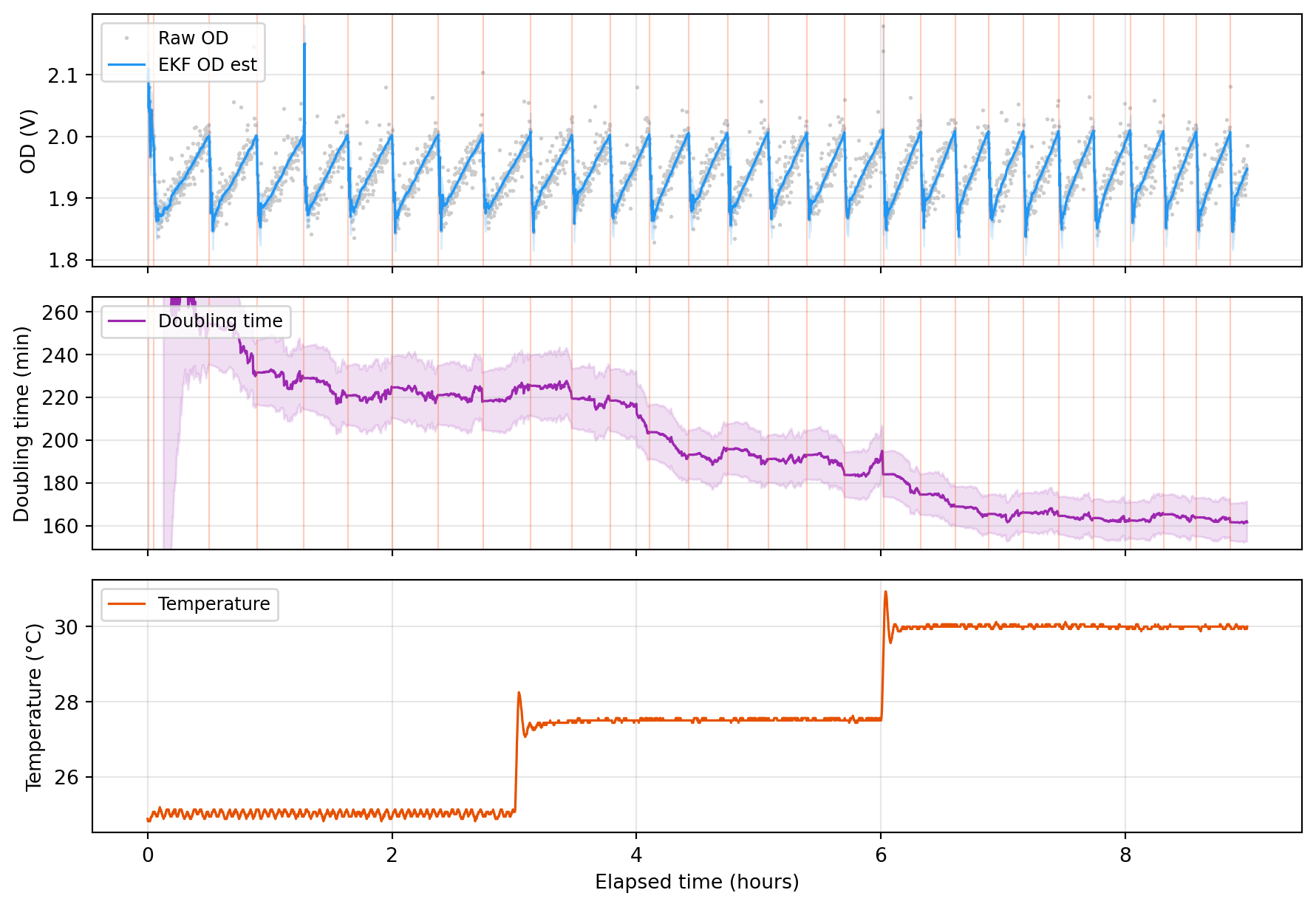
ax_od.plot(t_h, raw_od, '.', color='#cccccc', markersize=2, label='Raw OD', zorder=1)
ax_od.plot(t_h, ekf_od, '-', color='#2196F3', linewidth=1.2, label='EKF OD est', zorder=3)
ax_od.fill_between(t_h, ekf_od - od_sd, ekf_od + od_sd, color='#2196F3', alpha=0.15, zorder=2)
for pt in pump_times:
ax_od.axvline(pt, color='#FF5722', alpha=0.3, linewidth=0.8)
ax_od.set_ylabel('OD (V)')
ax_od.legend(loc='upper left', fontsize=9)
ax_od.grid(True, alpha=0.3)
dt_upper = np.where(np.isfinite(dt_min) & np.isfinite(dt_sd_min), dt_min + dt_sd_min, np.nan)
dt_lower = np.where(np.isfinite(dt_min) & np.isfinite(dt_sd_min),
np.maximum(dt_min - dt_sd_min, 0), np.nan)
ax_dt.plot(t_h, dt_min, '-', color='#9C27B0', linewidth=1.2, label='Doubling time')
ax_dt.fill_between(t_h, dt_lower, dt_upper, color='#9C27B0', alpha=0.15)
for pt in pump_times:
ax_dt.axvline(pt, color='#FF5722', alpha=0.3, linewidth=0.8)
finite_vals = dt_min[np.isfinite(dt_min)]
if len(finite_vals):
p5, p95 = np.percentile(finite_vals, [5, 95])
margin = (p95 - p5) * 0.15
ax_dt.set_ylim(max(0, p5 - margin), p95 + margin)
ax_dt.set_ylabel('Doubling time (min)')
ax_dt.legend(loc='upper left', fontsize=9)
ax_dt.grid(True, alpha=0.3)
ax_temp.plot(t_h, temp, '-', color='#E65100', linewidth=1.2, label='Temperature')
ax_temp.set_ylabel('Temperature (°C)')
ax_temp.set_xlabel('Elapsed time (hours)')
ax_temp.legend(loc='upper left', fontsize=9)
ax_temp.grid(True, alpha=0.3)
plt.show()